Initiation of protein synthesis:
Each prokaryotic ribosome, shown schematically in the figure, has three binding sites for transfer RNAs. The aminoacyl-tRNA binding site (or A site) is where, in during elongation, the incoming aminoacyl-tRNA binds. The peptidyl-tRNA binding site (or P site) is where the tRNA linked to the growing polypeptide chain is bound. The exit site (or E site) is a binding site for tRNA following its role in translation and prior to its release from the ribosome. All three sites (A, P and E) are formed through the rRNA molecules in the ribosome.
The first codon translated in all mRNAs is AUG that codes for methionine. This AUG is known the begin codon or initiation codon. Obviously, other AUG codons also occur internally in an mRNA where they encode methionine residues internal to the protein. Two varients tRNAs are used for these two kinds of AUG codon; transfer RNAfmet is used for the initiation codon and is known the fMet initiator tRNA whereas tRNAmMet is used for internal AUG codons. In the prokaryotes the first amino acid of a new protein is N-formylmethionine abbreviated Met fMet. Therefore the aminoacyl-tRNA used in initiation is fMet-tRNAf also abbreviated as fMet-tRNAfMet. It is essential which the correct AUG is used as the initiation codon because this groups the correct reading frame for translation. A short sequence rich in purines (5'-AGGAGGU-3'), known as the Shine-Dalgarno sequence, lies 5' to the AUG initiation codon described in the figure and is complementary to category of the 16S rRNA in the little ribosomal subunit. Thus this is the binding site for the 30S ribosomal subunit that then migrates in a 3' direction along the mRNA until it encounters the AUG initiation codon. Therefore the Shine-Dalgarno sequence delivers the ribosomal subunit to the correct AUG for initiation for translation.
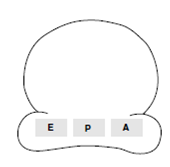
Figure: Schematic of a prokaryotic 70S ribosome showing the peptidyl-tRNA site (P site), aminoacyl-tRNA site (A site) and exit site (E site).
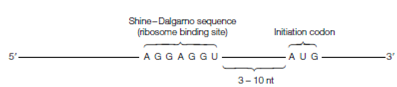
Figure: The Shine-Dalgarno sequence in prokaryotic mRNA.
Initiation of protein synthesis needs proteins known as initiation factors (IFs). In the prokaryotes, three initiation factors are IF-1 and IF-2 and IF-3 essential. Because of the complexity of the procedure, the exact order of binding of IF-1 and IF-2, and IF-3 and fMet-tRNAfMet and mRNA is still unclear. One present model is shown in Figure and is described below.
- Initiation starts with the binding of IF-1 and IF-3 to the small (30S) ribosomal subunit. Their role is to end the 30S subunit binding to the 50S subunit in the absence of mRNA and fMet-tRNAfMet functional ribosome. Which would result in a non-
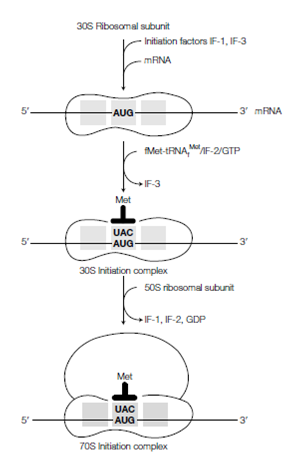
Figure: Initiation of protein synthesis in prokaryotic cells.
- The small subunit then binds to the mRNA through the Shine-Dalgarno sequence and moves 3' along the mRNA until it situates the AUG initiation codon.
- The initiator transfer RNA charged with N-formylmethionine and in a complex with IF-2 and GTP (fMet-tRNAfMet/IF-2/GTP) now binds.
- IF-3 is released.
- The difficult of mRNA, fMet-tRNAf Met, IF-1, IF-2 and the 30S ribosomal subunit is known the 30S initiation complex.
- The huge (50S) ribosomal subunit now binds with the release of IF-1 and IF-2 and hydrolysis of GTP, to appearance a 70S initiation complex.
One most important point to note is which, unlike all other aminoacyl-tRNA molecules (that bind to the A site), the binding of fMet-tRNAf Met occurs straightly into the P site.